Watch tutorials, webinars and informative videos about PreSens optical sensor systems.
Evaluating the SDR SensorDish® Reader for Strain Development
Monitoring different E. coli strains inside deep well OxoDishes®
Olaf Christensen
Lonza Ltd., Visp, Switzerland
We conducted several tests to evaluate the applicability of the SDR SensorDish® Reader combined with deep well OxoDishes® in strain development. We measured oxygen consumption of induced and non-induced E. coli strains with the SDR, and conducted media tests changing the substrate in cultures producing different proteins. Oxygen measurements recorded in parallel experiments showed excellent reproducibility, and also enabled us to get detailed information on the metabolic status of the cultures at all times. Compared to standard end-point analyses, this real-time monitoring system offers new options, e. g. when developing harvesting strategies and optimizing media.
Oxygen is a vital parameter in microbial and cell culture. Monitoring O2 during production processes helps avoid failure due to oxygen limitation and gives valuable information about the metabolic status of the cultures. The use of multiwell plates with integrated oxygen sensors at the bottom of each well - like the OxoDishes® by PreSens - is very interesting for strain development studies, as parallel online monitoring of multiple strains can be conducted in one experimental run. The deep well version of the OxoDishes® allows shaking the cultures. A Duetz clamp system and metal covers are used to fix the SDR SensorDish® Reader with the deep well plate on top inside the shaker. We tested if this system is suitable for strain development studies, and if the online oxygen readings can contribute to identifying promising strains.
E. coli Culture in Deep Well OxoDishes®
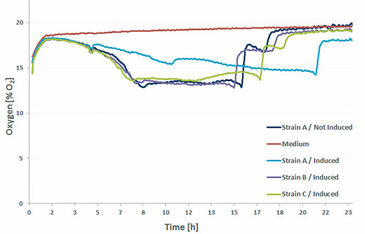
Three different E. coli strains - all producing the same protein - were cultured in a deep well OxoDish® with a volume of 2 mL/well, at 30 °C and 280 rpm (d = 50 mm). All three strains were induced; additionally oxygen in the not induced Strain A and medium only (control) was measured (Fig.1). The decreasing oxygen values during the first 8 h indicate the exponential growth and high metabolic activity of all strains. In the next phase growth stopped and oxygen ingress in the medium equaled oxygen consumption by the bacteria, showing in stable oxygen values of about 14 % O2 for the next 8 h. Then substrate limitation set in and oxygen levels started to rise again due to stopped metabolic activity and cell death. A small plateau in the oxygen curves after 15 h of cultivation indicates that a metabolic shift took place and the E. coli cultures switched to metabolizing another substrate. After 1.5 h this substrate had also been consumed and oxygen values rose to air saturation. One exception is the induced Strain A which showed unusual behavior throughout the experiments. It grew slower, so substrate limitations set in a lot later than in the other cultures. One explanation might be the extremely high production rates of Strain A so the bacteria might have been stressed by the final product.
Media Tests with the SDR
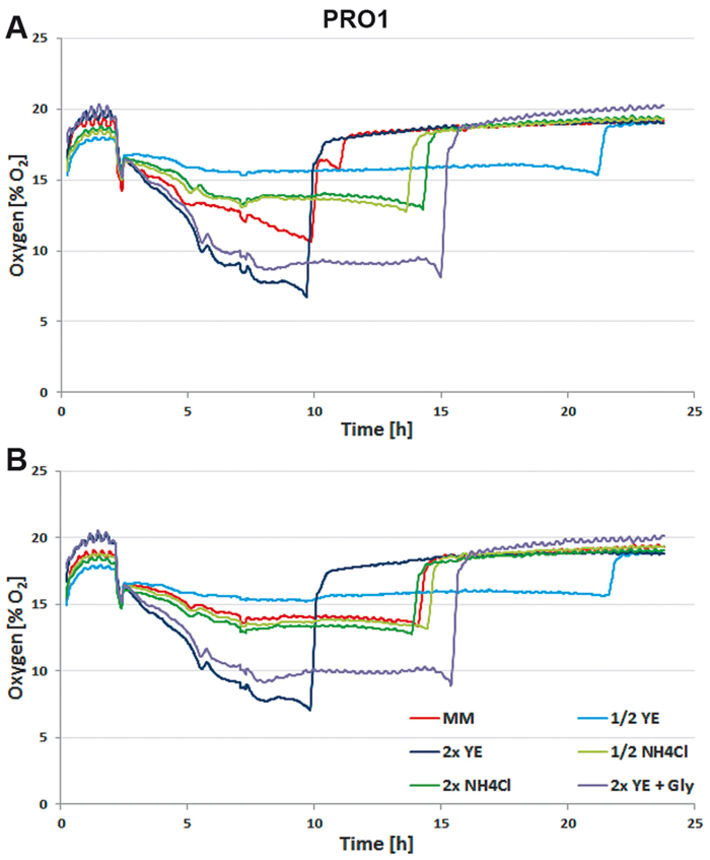
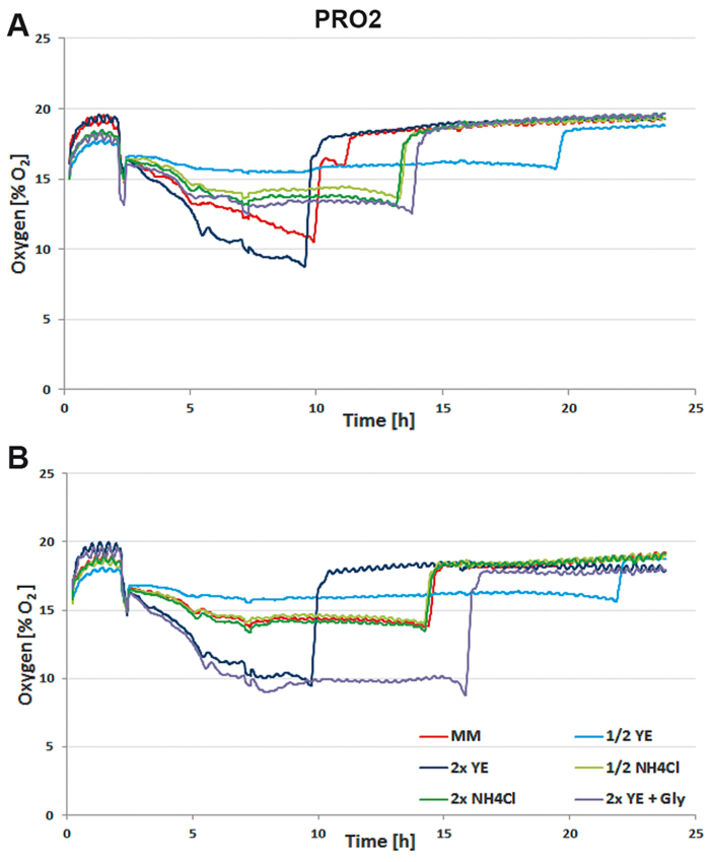
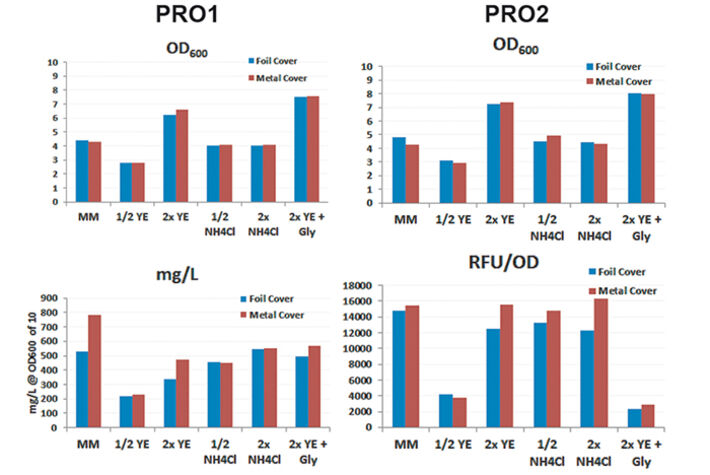
For media tests induced or not induced E. coli Strain C producing either protein 1 (= PRO1, Fig.2) or protein 2 (= PRO2, Fig. 3) were cultured in deep well OxoDishes® at the same conditions as mentioned above and either covered with a foil membrane (to avoid evaporation) or the metal covers of the clamp system. Different media compositions were used to evaluate culture performance and production when substrates are changed. We used semi-defined medium (MM) (Lonza AG, Switzerland), medium with half or double the amount of yeast extract (1/2 or 2x YE), half or double the amount of ammonium (1/2 or 2x NH4Cl), and double the amount of yeast extract and glycerine (2x YE + Gly). Oxygen measurements taken over a period of 24 hours showed that independent from the product the data of E. coli in the different media are very well comparable. While cultures with double the amount of YE showed fastest growth, the substrate was also consumed fastest. Double the amount of YE + Gly also showed fast growth, but the additional C-source glycerin extended the activity phase of the culture. Double or half the amount of NH4Cl did not seem to affect growth as cultures showed the same metabolic activity in both media, so NH4Cl is not limiting. Cultures with half the amount of YE grew slowest, reaching substrate limitation only after more than 20 hours. These results show, the more YE is added to the medium, the faster the E. coli cultures grow, independently from the product, and glycerin can be used to extend the activity phase. Comparing not induced and induced cultures the oxygen readings show that only the induced PRO1-producing culture in MM is growing slower and has a longer activity phase than the non-induced. In PRO2-producing E. coli the induced cultures have longer activity phases, reaching limitation later than the not induced cultures. This might be due to a metabolic shift and subsequent consumption of another substrate. OD and expression analyses (Fig. 4) were in good accordance with the online measurements. Here cultures in 2x YE also exhibited strongest while those in 1/2 YE showed weakest growth, and both media with different NH4Cl concentrations as well as cultures in minimal medium produced similar OD. The use of standard foil covers compared to the metal covers had no effect on productivity, and there was no difference in PRO1 expression using a standard deep well plate and the deep well OxoDish® (data not shown).
Conclusion
The SDR SensorDish® Reader in combination with the deep well OxoDishes® is easy to set up and gives excellent reproducibility in oxygen readings. We were able to compare culture performance of different strains, and showed that media tests can be performed with the SDR comparing different components in one experimental run. Altogether this system offers a huge advantage compared to commonly applied end-point analyses.
Lonza Ltd makes no warranty as to its accuracy or completeness of the above evaluation and Lonza Ltd assumes no obligation or liability in this respect. All trademarks belong to their respective owners and are used here only for informational purposes.